Analyzing Standards (GC)
The final step in evaluating the system performance prior to analyzing samples is to analyze a suitable standard(s)
Stable isotope standards
It is important that consideration must be given to the types of standards used to assess the instrument prior to (and during) sample analysis. The principle of identical treatment of the standards and samples should be followed as closely as possible in that both should be analyzed in the same way. Ideally, the standards should contain the same compounds (or compound classes) to the samples, have stable isotope values that span the expected range and should be analyzed using the same GC conditions and in similar concentrations to the samples.
Analyzing a GC standard with known isotopic compositions can serve to evaluate chromatography, sensitivity, linearity, accuracy and also provide a mechanism to calibrate sample measurements. A vast array of stable isotope standard materials are readily available from the International Atomic Energy Agency (IAEA), National Institute of Standards and Technology (NIST) and the United States Geological Survey (USGS), however these are typically single compounds and often not GC amenable. For GC standards mixes we would recommend the standards produced by Arndt Schimmelmann from Indiana University.
A cost effective approach is to also produce in-house standard mixes; here the objective is to obtain a suite of pure, isotopically homogeneous compounds with varying values that have been calibrated using the IAEA/NIST/USGS standards. This can be achieved by analyzing the potential in-house standard components via EA IRMS and then creating a suitable mixture for GC analysis.
GC methods
To analyze the standard(s) there are two methods that are intrinsically linked that need to be considered, the IonOS method and a 7890 GC method.
The default IonOS GC method is called ‘GC Analysis’ and can be accessed in same way as the ‘stability’ and ‘linearity’ methods discussed above, i.e. Under ‘tasks’ tab, select ‘methods’ in the ‘manage’ section.
The input parameters for ‘GC analysis’ as shown in Figure 6-45 illustrate that the default GC method (the 7890 GC method) is the one currently loaded into the GC, there is a 4 minute solvent delay (solvent diverted away from the IRMS), no extra heart cuts (diverting GC flow away from the IRMS), the autosampler and tray are to be used and the sample is in vial position 1, the injection volume will be 1ul and there will be three monitoring gas pulses at the beginning of the analysis and a further three at the end when the 7890 GC method has ended. Similar to before these parameters can be adjusted within the task list if the ‘editable in the task list’ checkbox has been ticked.
A ‘7890 GC method’ can be either created on the GC keypad or directly through IonOS. Figure 6-46 shows the 7890 GC ribbon tab and under method files there is an option to ‘download (a method) from the GC, ‘upload to the GC’ or ‘manage’ the method files.
Using the GC keypad
To program the GC via the keypad please follow the instructions below, ensure the parameters are altered to suit the particular analysis and column (i.e. don’t exceed the maximum temperature and ensure the dimensions of the column are input correctly). For more comprehensive details on how to program the GC, please refer to the Agilent 7890 GC manual.
In the following example, the GC method is setup in split mode (split ratio 10) using a 30m GC column with 250um ID and 0.25um film thickness.
Configuration
Select the ‘Configuration’ button
Select ‘Front Inlet’ (if that is the inlet that is being used)
- Set ‘Equilibrium’ time to ‘1’
- Set the ‘Detector’ to ‘Back Detector’ (the FID is in the back position)
Select ‘Column’ and press ‘1’ (the GC can store details on up to 6 columns)
- Set ‘Length’ to ‘30m’
- Set ‘Diameter’ to ‘250um’
- Set ‘Film thickness’ to ‘0.25um’
Setting the temperatures and gas flows
Select the ‘Oven’ button and scroll through the options using the arrows checking / changing the following parameters:
- Set ‘Temp’ to ‘50’ (initial temperature)
- Set ‘Initial time’ to ‘1’ (isothermal hold time in min)
- Set ‘Ramp’ to ‘10’ (temperature ramp rate of 10C /min)
- Set ‘Final temp’ to ‘300’ (final temperature)
- Set ‘Final time’ to ‘10’ (isothermal hold time at the final temperature)
(Ensure that the final temperature does not exceed the maximum temperature allowed by the GC column)
Select the ‘Front Inlet’ button and scroll through the options using the arrows checking / changing the following parameters:
- Set ‘Temp’ to ‘270’ (injector temperature)
- Set ‘Split’ to ‘10’ (split ratio set to 10)
(if you want to analyze in splitless mode, then select this after pressing the ‘mode’ button the keypad)
Select the ‘Column’ button and scroll through the options using the arrows checking / changing the following parameters:
- Select ‘constant flow’
- Set ‘Flow’ to 1.0 ml/min
Select the ‘Back Det’ button and scroll through the options using the arrows checking / changing the following parameters:
- Set ‘Temp’ to 300
- Press ‘Flame on’ (once the temperature is reached, air and hydrogen will mix and will be ignited into a flame for the FID)
(if you do not want to include the FID as part of the method, then do not switch it on)
Select the ‘Front Injector’ button if the autosampler is being used and scroll through the options using the arrows checking / changing the following parameters:
- Set ‘Solvent A pre wash’ to ‘3’ (cleans the syringe three times with solvent from wash bottle A prior to analysis)
- Set ‘Solvent A post wash’ to ‘3’ (cleans the syringe three times with solvent from wash bottle A after the analysis)
- Set ‘Solvent B pre wash’ to ‘3’ (cleans the syringe three times with solvent from wash bottle B prior to analysis)
- Set ‘Solvent B post wash’ to ‘3’ (cleans the syringe three times with solvent from wash bottle B after the analysis)
- Set ‘Sample washes’ to ‘zero’ (A ‘sample wash’ is not recommended as sample is taken up into the syringe during the washing process and discarded to waste, therefore only useful if sample volume is abundant)
- Set ‘Sample pump’ to ‘2’ or ‘3’ (This is the recommended alternative to the sample wash)
(Most parameters can be kept as default and the injection volume will be overwritten by the software)
Using IonOS
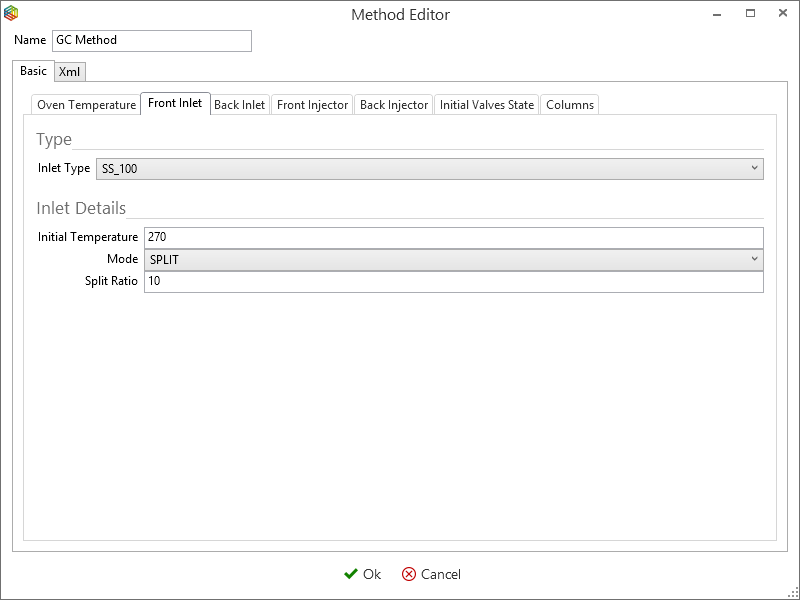
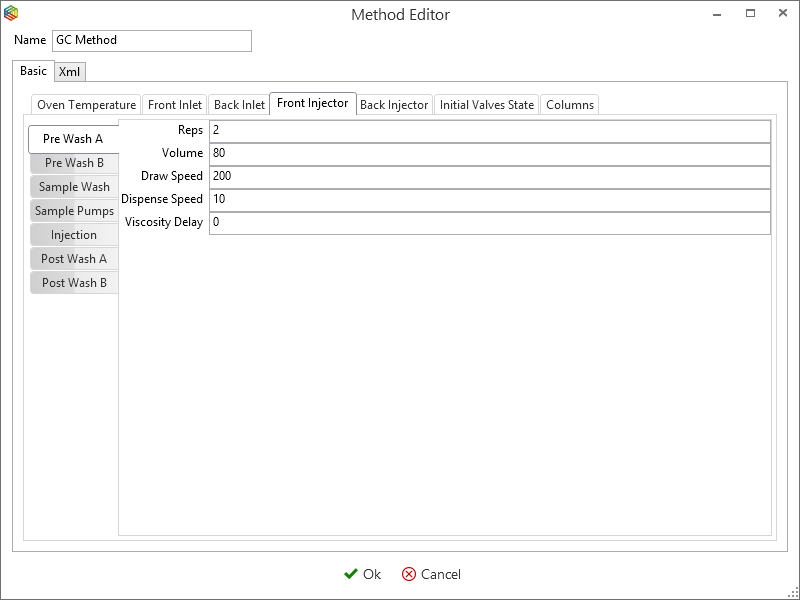
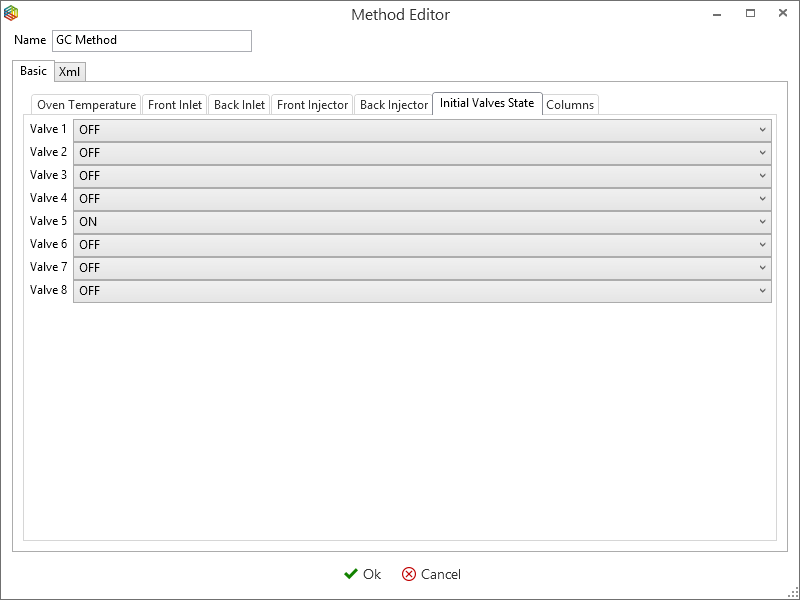
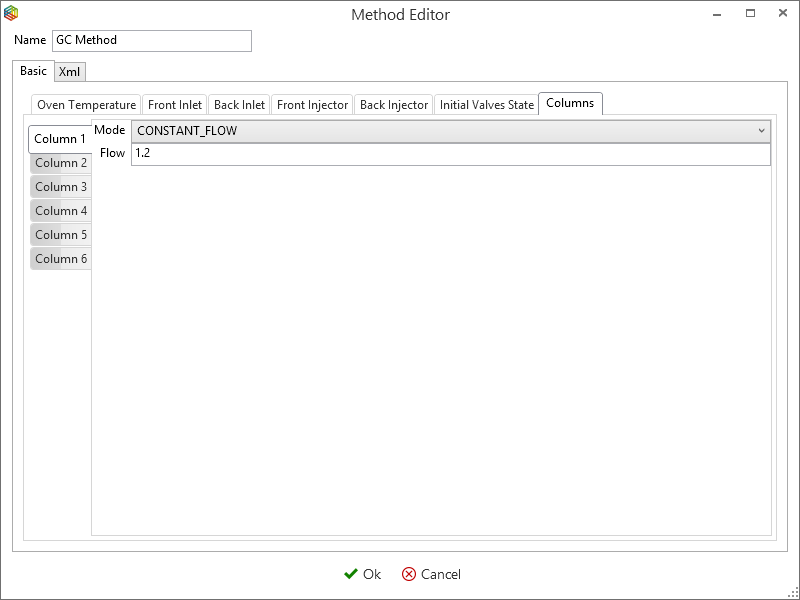
Although you can program the 7890 GC method entirely within IonOS, it is easier to copy and modify an existing method. Select ‘manage’ under the 7890 GC tab (Figure 6-46) to open the ‘GC methods global settings’ window (Figure 6-48). This contains a list of 7890 GC methods that are stored within IonOS, here you can choose to add, copy, delete and modify them. By selecting a method, e.g. ‘GC method’ and pressing the edit icon (Figure 6-47), the Oven temperature (Figure 6-48), Front Inlet (Figure 6-49), Front Injector (Figure 6-50), Initial Valve States (Figure 6-51) and the Columns (Figure 6-52) settings can be configured.
Running the standard
After the GC methods have been established, the analysis is ready to proceed. This is achieved by running a task with the ‘GC analysis’ method selected in the task list. Although all the input parameters are pre-determined within the method, these can be altered within the task list. In the above example, the option of using the ‘autosampler' (with further options of using the ‘tray’ or ‘turret’), the ‘vial number’ and the ‘solvent delay’ time can all be overwritten within the task list.
As it is important to avoid solvent traveling through the furnace tube towards the IRMS, it is advisable when analyzing a standard for the first time to divert all the GC flow towards the FID to determine the solvent elution time. To view the FID chromatogram, see the ‘Data Analysis’ window under the ‘FID’ tab. Once the solvent peak has been identified, the ‘solvent delay’ time in the method or task list should be set to greater than this.
A quick way to create an ‘FID only’ run is to adjust the ‘solvent delay’ in the task list to the length of the GC analysis time, e.g. 36 minutes in the above example (’00:36:00’).
Autosampler injection
When setting up an autorun using the Agilent 7693 liquid autosampler, it is important the ‘Use Autosampler’ setting is set to ‘true’ (this can be done in the method or in the task list) and that, depending on the instruments configuration, ‘tray’ or ‘turret’ is selected under the ‘injector’ column (Figure 6-53). Ensure that the ‘vial number’ corresponds to where the sample / standard is positioned in the autosampler tray or turret and the correct GC method is selected.
To begin analysis, highlight the task(s) and then press ‘start’ in the software.
When evaluating the system with a standard it is advisable to analyze it several times to determine the level of precision that can be achieved. To investigate linearity, inject different concentrations of the same standard or inject different volumes by altering the injection volume parameter in the task list; be careful that the vapour volume does not to exceed the volume of the injector liner (Table 4-2).
Manual injection
To carry out a manual injection, the ‘Use Autosampler’ parameter should be set to ‘False’ in the method or task list (Figure 6-54). When altered in the task list, the parameters associated with the autosampler, i.e. the ‘injector’ and ‘vial number’, will be ignored.
To begin the analysis, highlight the task(s) and press ‘start’ in the software; when the GC is in a read state, the system status indicator will turn amber and a message stating that the ‘GC is ready for injection’ will appear in the system status window. At this stage the sample can be injected manually with a syringe, followed immediately by the pressing of the start button on the GC keypad.
One slight variation to the procedure is when the GC method is setup for a splitless injection using the S/SS injector. During a splitless injection, the split vent is required to be shut just prior to injection (and for a period of between 0.75 and 1 minute after); this is achieved automatically when using an autosampler, however for a manual injection method this vent is required to be shut manually by the analyst. When the system status indicator is amber and the message stating that the ‘GC is ready for injection’ is visible in the status window, the user is required to press the ‘prep Run’ button on the GC keypad before injecting.
Evaluation of Data
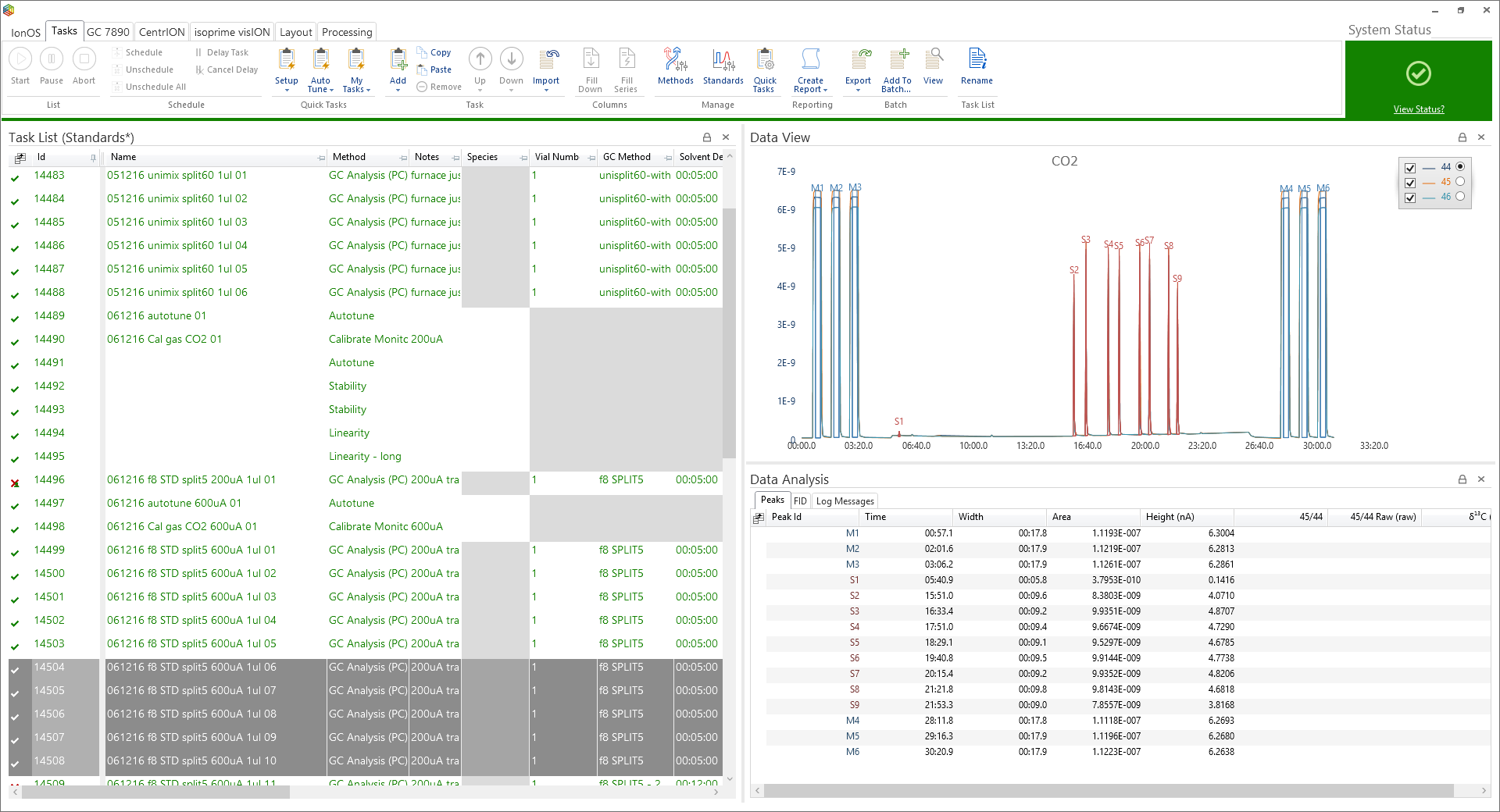
Within the system page the raw data and the chromatogram from each run can be seen in the ‘data analysis’ and ‘data view’ windows, respectively, however to collectively view (and process) all the runs together it is advisable to group them within a batch.
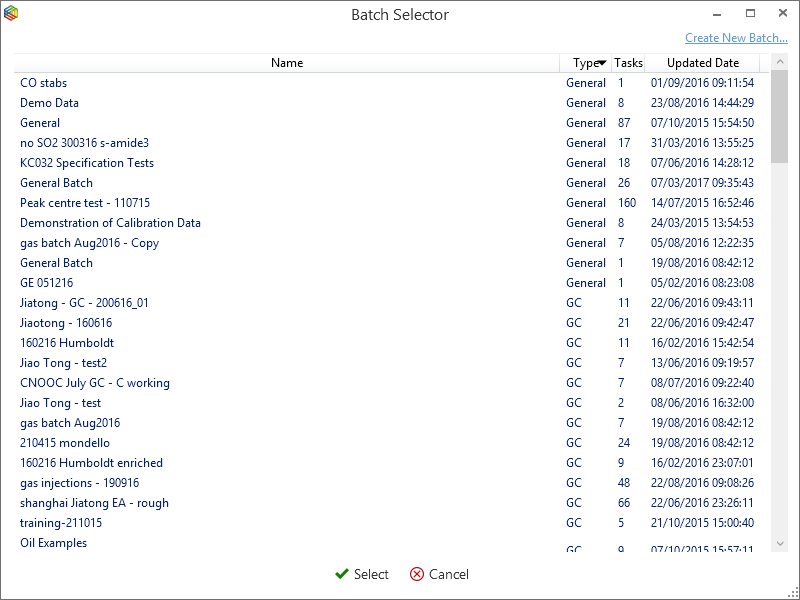
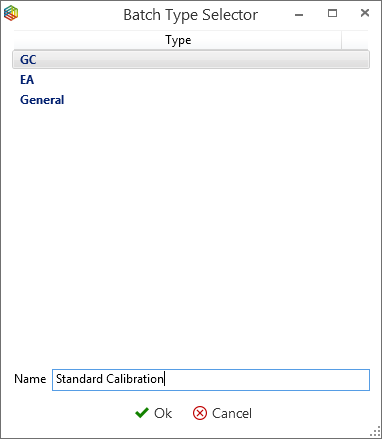
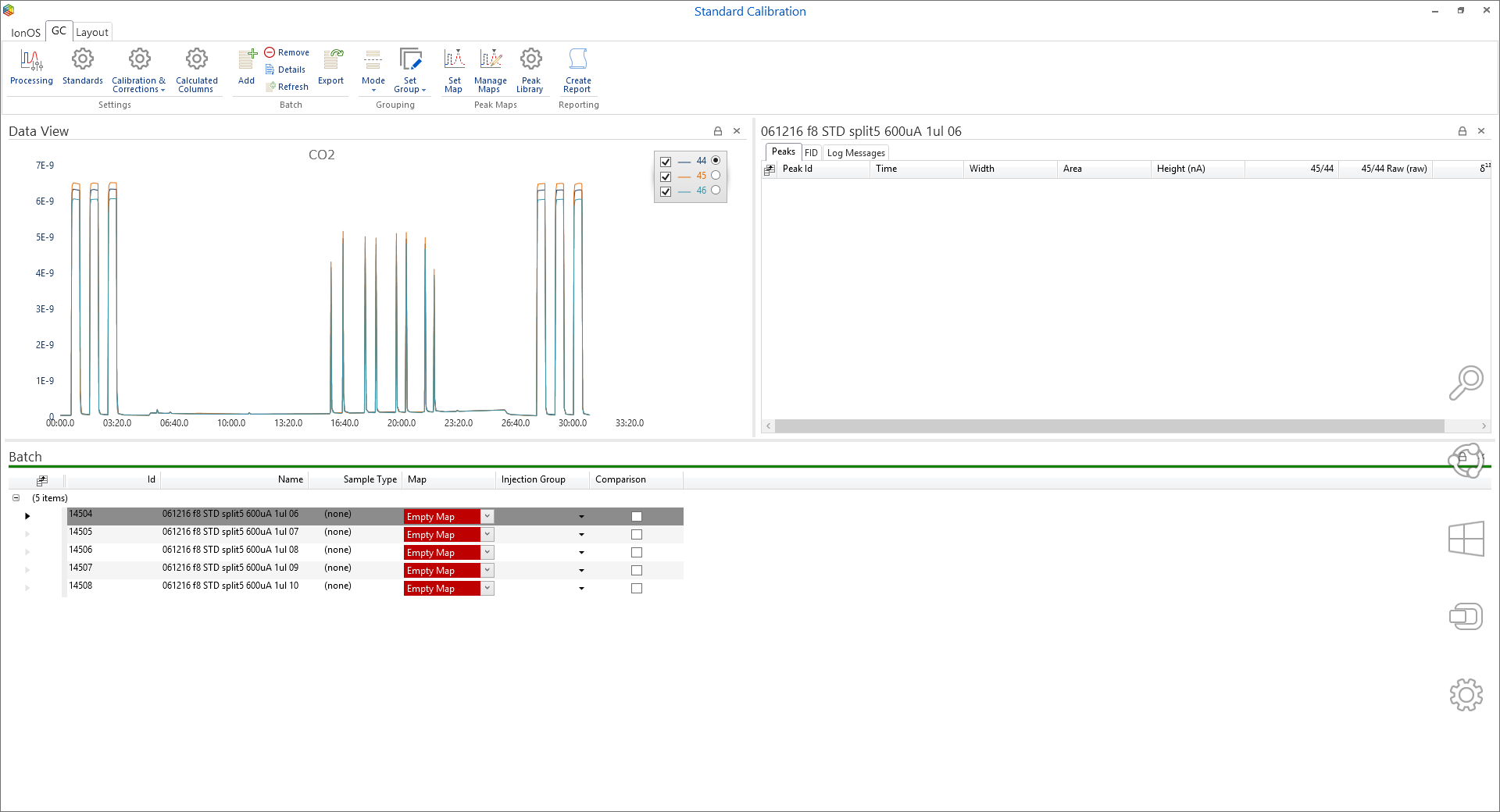
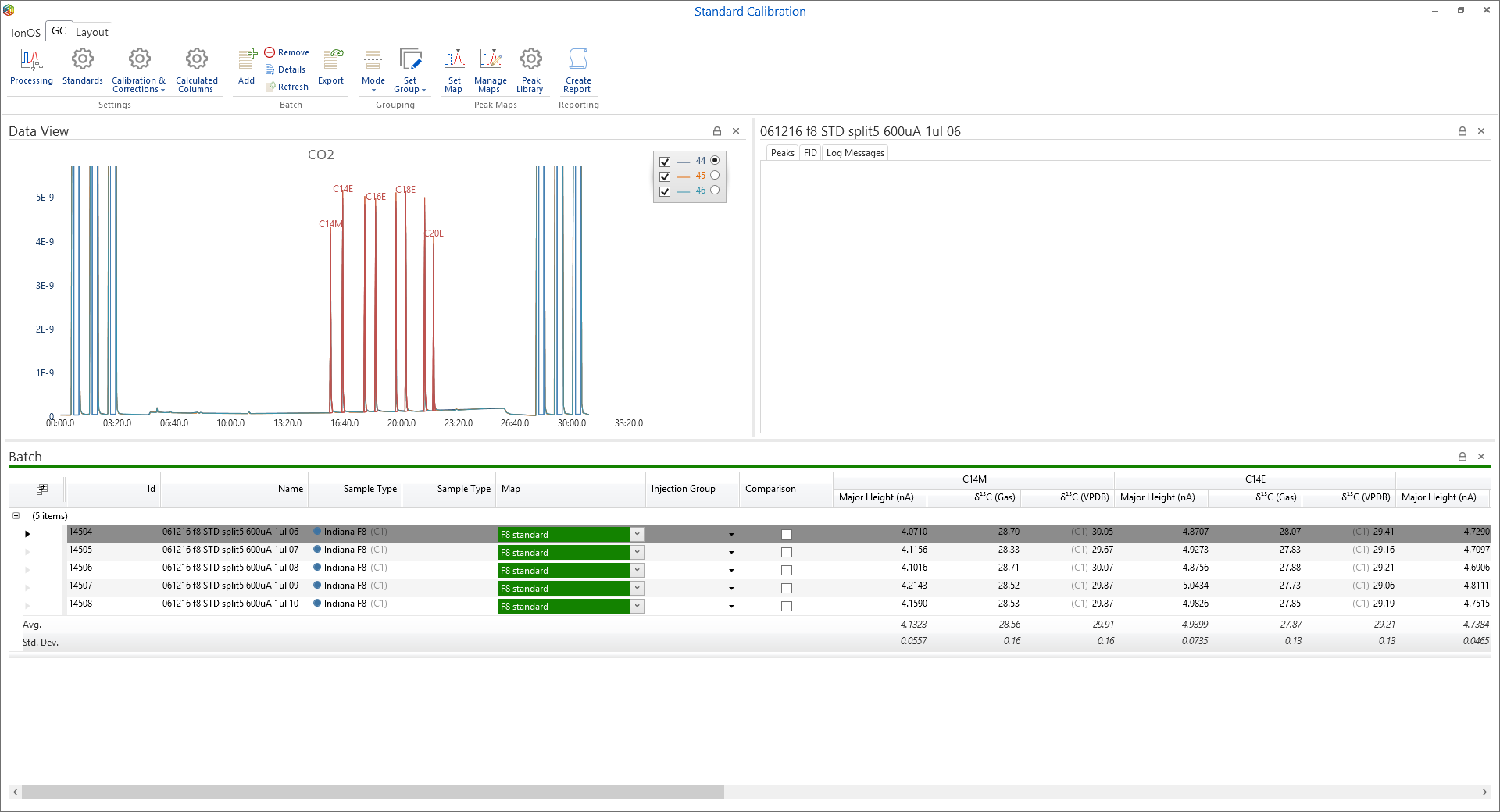
To add task runs to a batch, simply highlight the runs and under the ‘Tasks’ tab, in the ‘Batch’ section, click on the ‘Add to Batch’ icon (Figure 6-55) where the ‘batch selector’ window will open (Figure 6-56). Within ‘batch selector’ there is an option to add the tasks to pre-existing batches or to create a new one by clicking on the ‘create new batch’ option in the top right corner. When creating a new batch, the user is prompted to select what type, i.e. ‘GC’, ‘EA’ or ‘General’ and create a name for the batch (Figure 6-57). The ‘GC’ and ‘EA’ batches are for processing GC and EA data, while a ‘general’ batch is for grouping stability, linearity, background scans, etc. Figure 6-58 displays a GC batch called ‘standard calibration’, which contains five GC runs. Further runs can be added from the task list like before or by pressing the ‘Add’ batch icon under the ‘GC’ tab.
The task runs that were added to the batch were all standards and these can be assigned as such within the batch to enable automatic standard compound identification; here the software looks for the unique signature of isotopic results associated with the standard to correctly identify the compounds and when combined with the peak mapping functionality the individual peaks for all the standard runs are correctly labeled.
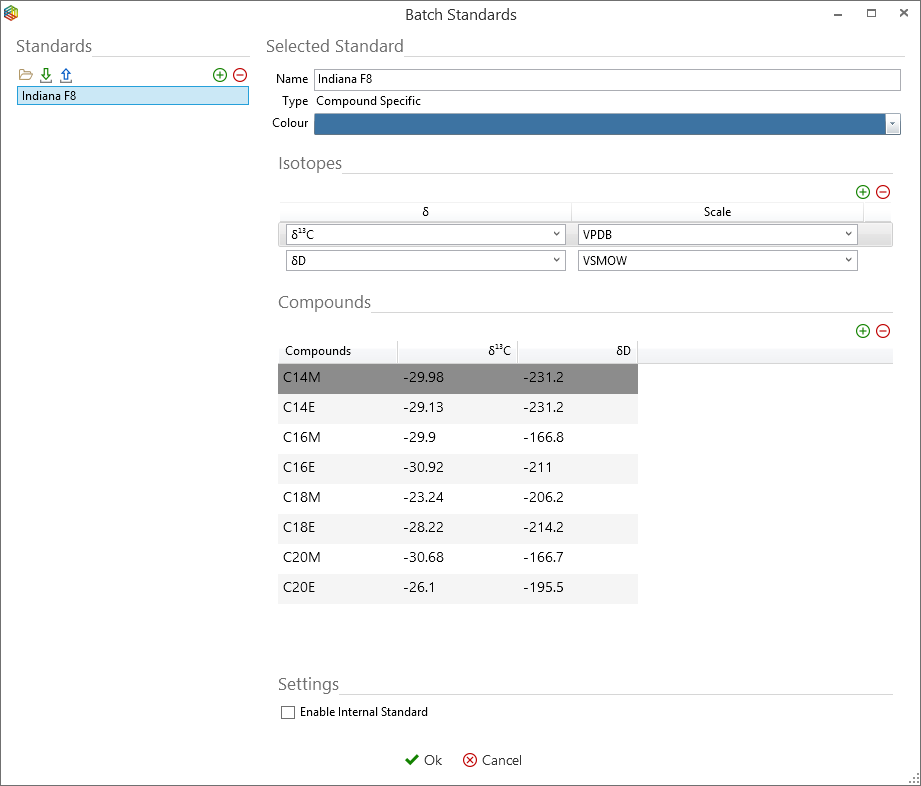
First the standard data, i.e. peak labels and known isotopic values are required to be made available to the batch by pressing the ‘standards’ icon under the ‘GC’ tab. If no standards are listed then they can be included by pressing the ‘+’ icon and typing in all the peak compounds present and their corresponding stable isotope values. However, it is recommend to store the standard data within the global settings of IonOS and these can be simply imported by pressing the ‘import global’ icon (Figure 6-59 and Figure 6-60). To create or import standards into the global settings, please go to the IonOS Home page and under IonOS, select settings > global settings > Processing > standards.
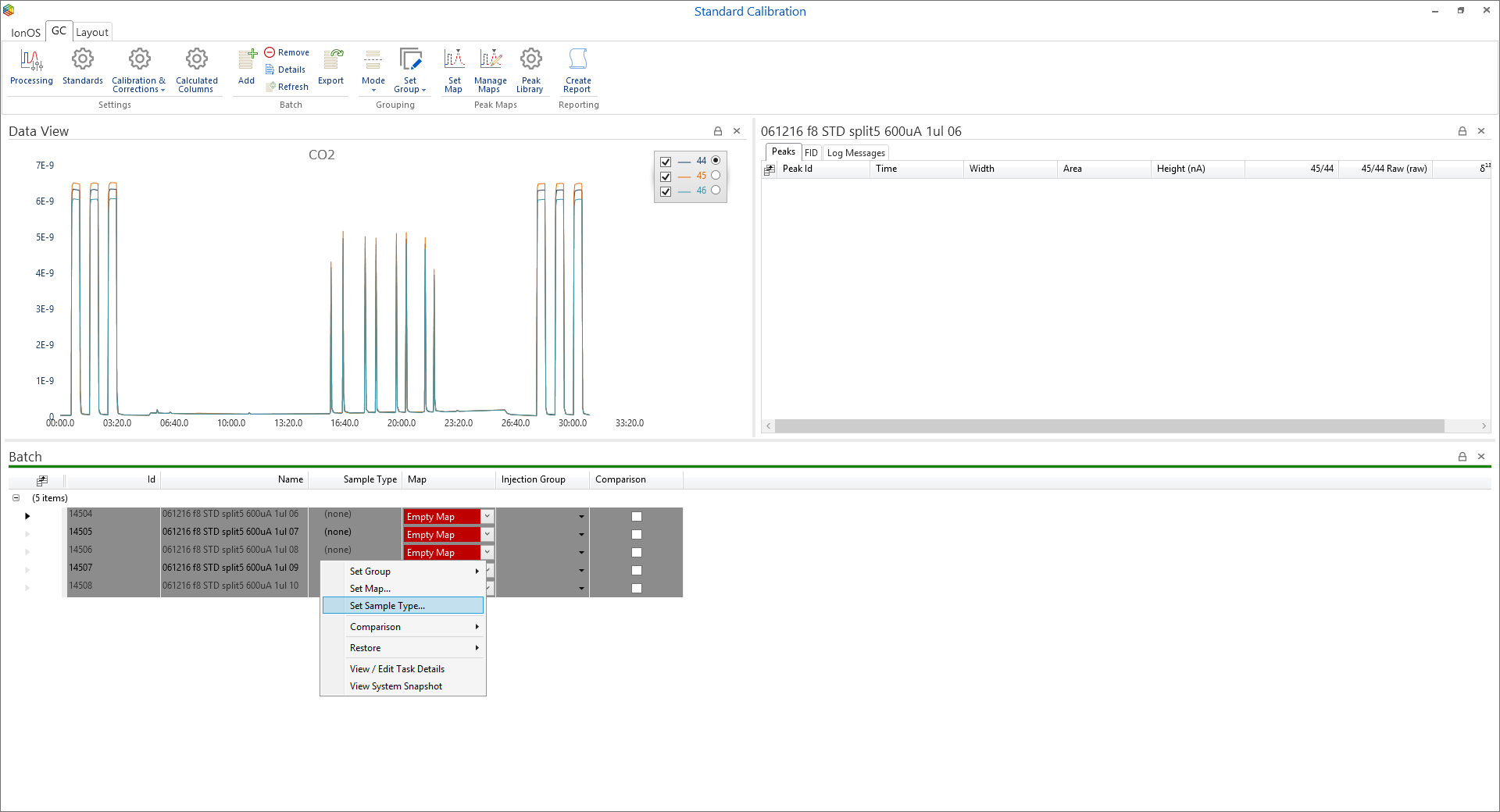
Once the required standard is available within the batch then each standard run is assigned as that standard. This done by highlighting the required runs, right-clicking and selecting ‘Set Sample Type…’ from the menu (Figure 6-61); the ‘sample type’ options will open and the standard can be selected (Figure 6-62).
Peak mapping is a process to select and identify target components within one chromatogram and to apply this across multiple injections; this time saving functionality is particularly useful in processing complex chromatograms. When a run has been identified as a standard, the process of identification of the target components can be done automatically using the Automatic Compound Identification feature where the unique isotopic signatures enable correct peak identification .
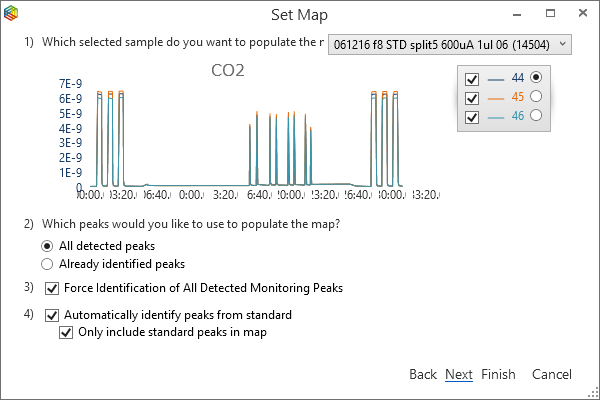
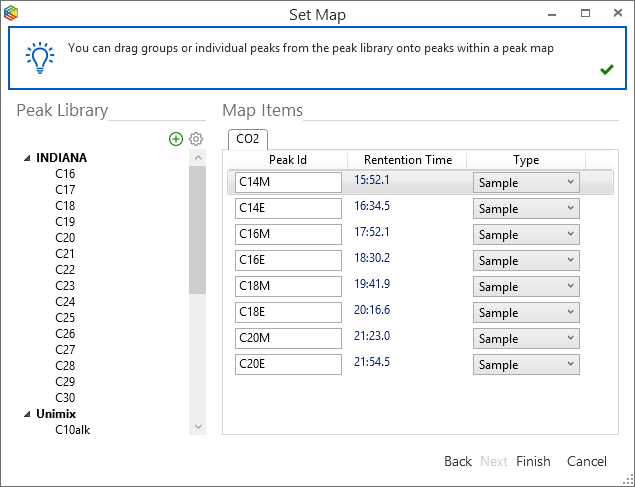
By highlighting the standard runs in the batch, select ‘Set Map’ from the menu to open up the ‘Set Map’ window (Figure 6-63). There are options to ‘Create a new map’, ‘Assign an existing map’ or ‘Assign an empty map’; select ‘Create a new map’ and for the ‘peak identification’, choose the ‘Let IonOS automatically identify the peaks’ followed by ‘Next’ to reveal more options (Figure 6-64). Select which sample to base the map on from the dropdown list in the top right-hand corner, the chromatogram will also be shown in the dialogue. Keep the default IonOS settings of ‘All detected peaks’ when creating the peak map and to ‘Force identification of all detected monitoring peaks’. To enable ‘automatic compound identification’, ensure that the checkboxes ‘Automatically identify peaks from standard’ and ‘only include standard peaks in map’ are ticked. Pressing ‘Next’ reveals the generated peak map that includes peak labels, which were assigned from the standard, and retention times (Figure 6-65). By pressing ‘finish’, the peaks are labeled in the chromatograms and their stable isotope values relative to the monitoring gas are displayed (Figure 6-66).
The averages and standard deviations across the standard runs are calculated automatically within the batch (Figure 6-66), however to further evaluate the quality of the data we can compare the measured stable isotope values for the individual components with their expected values using the ‘Calibration & Corrections wizard’ (Figure 6-67).
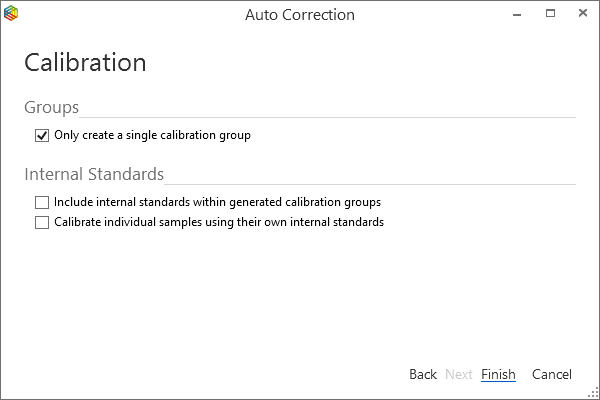
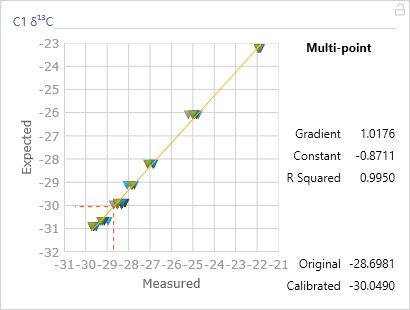
Grouping the standards into a single calibration group (Figure 6-68) and pressing ‘finish’ will automatically compare, in this case, the d13C value relative to the monitoring gas for each component with its expected value. Figure 6-69 displays the calibrated data and by hovering the cursor over ‘(C1)’ the calibration curve can be displayed along with the gradient of the line, the intercept (or constant) and if there are more than two points, the R squared value (statistical measure of how close the data are to the line)(Figure 6-70). Ideally the gradient should range from 0.95 to 1.05 and the R squared value greater than 0.99.
To establish if the system is ready to analyze samples is very much dependent on the analyst and their specific application. If the answer to the following questions is yes, then the instrument is ready to analyze samples:
- Is the level of sensitivity good enough for the samples / application?
- Is the precision good enough?
- Is the linearity good enough?
- Is the calibration curve as expected?
If not, please refer to Troubleshooting.